<p>Small Bugs, Big Data</p>
<p><span class="Smaller90">Inaugural lecture for the Chair of Clinical Microbiology at Radboud University<br /></span><span class="Smaller90">17 January 2017 - </span><span class="Smaller90">Radboud University, Nijmegen</span></p>
Introductory remarks
Mijnheer de rector magnificus,
leden van het college van bestuur van de Radboud Universiteit,
leden van de raad van bestuur Radboudumc,
collega’s, familie, vrienden en andere aanwezigen,
collaegues, collaborators and friends abroad watching by live stream,
It is a great honour to stand here and give my inaugural lecture for the Chair of Clinical Microbiology at Radboud University. It is also an honour to do this in the presence of the invisible 14 quadrillion microorganisms that you all collectively have brought into this room today; weighing about 700 kilograms in total. Each one of you has more microbial cells than you have human cells. And the genes of these microbes contribute more to the proper functioning of your body than your own genes.
Unravelling the human microbiota is one of the many scientific advances we are experiencing today. The exponential speed of innovations and subsequent changes will have a major impact on healthcare. We need to build a vision of how the newly-acquired knowledge and skills can be utilised in everyone’s best interest – from healthcare workers to patients. During this lecture, I will provide a snapshot of what I think these advances in our digitising world mean for clinical microbiology and of my own interest in the global problem of antimicrobial resistance. These advances have one thing in common: they produce and analyse lots of data on many small bugs.
Preparing for this lecture was a valuable experience as it encouraged me to reflect on my professional life. The experience of working and living with my family for nine years in Vietnam and transitioning to Radboudumc just over a year ago made me more conscious of the extreme differences we encounter in our world. From a country with a population of 95 million people, to the Netherlands, a country of 17 million people, where the number of clinical microbiologists in the province of Gelderland alone exceeds the total number of clinical microbiologists in Vietnam, a country with a higher infectious disease burden, widespread antimicrobial resistance, and a greater need for clinical microbiology.
Back in the Netherlands, I was struck by the high concentration of and access to expertise and skills and the multidisciplinary approach to health. This is unique and precious: it explains the success we have as a country in providing high-quality healthcare, in controlling infectious diseases and drug resistance, and in our world-class scientific pursuits. The Netherlands has a lot to offer in the struggle to control infectious diseases, starting from the early pioneers Antoni van Leeuwenhoek, who discovered bacteria, and Martinus Beijerink (1851-1931), who discovered viruses, to the cutting edge research conducted today at our and other institutions.
The Dutch love to explore; I experienced this myself as a child when my parents decided to move to Panama for several years. In retrospect, it has become clear to me that my family’s move to Panama shaped me in such a profound way that clinical microbiology was destined to become my career choice.
Panama and infectious diseases
In Panama, the Bridge of the Americas spans the Panama Canal, connecting North America with South America. It was built over a hundred years ago. My teenage years in Panama were my first encounter with the tropics and the dangers of mysterious tropical diseases. Before our move to Panama, we had to consult a travel clinic, where we received our vaccinations against hepatitis and yellow fever, and were given maps showing the locations of infectious diseases like malaria. I found this very mysterious and exciting: a whole invisible natural world was out there with its own rules and with the potential to cause serious harm to humans. My curiosity was further fed by my mother, who was reading the historical book The Path Between the Seas about the creation of the Panama Canal (see figure 1).
Figure 1. The path between the seas

She told me fascinating stories from that period. For instance, because so many canal workers became sick or died from yellow fever or malaria, completing the canal was considered almost impossible. The role of mosquitoes in transmitting these infections was being debated at the time. The scientists who suggested that mosquitoes where responsible and that large-scale vector control programmes were needed for disease control were initially ridiculed. One small detail I remember clearly from her stories was about ants in a Canal Zone hospital. Thousands of ants were crawling all over the helpless patients in their beds. They decided pragmatically to put the bed legs into pans of water to prevent the ants from getting into the beds. As you can imagine, these pans with water were hardly ever refreshed, and the stagnant water became a breeding ground for mosquitoes. If you came to the hospital with a broken leg, you were likely to get a mosquito-borne disease on top of it.
In the end, the good news was that the mosquito hypothesis was finally accepted, vector control programmes were started and the canal was completed. This interplay between human behaviour and the microbial environment intrigued me. When I learned about the work of clinical microbiologists after my medical degree, I knew instantly that this was going to be my speciality. It provided boundless potential to combine infectious disease diagnostics with clinical consultation, infection prevention, research, public health, policy development and a multidisciplinary approach, all within a global context.
Atlas of Human Infectious Diseases
During my residency in clinical microbiology in Rotterdam, I enjoyed studying infectious disease maps, wondering how these were made, why diseases are where they are, and where the map-makers got their data. The idea behind the Infectious Disease Atlas project was to create a single resource illustrating the distribution of infectious diseases across the globe (see figure 2).
Figure 2. Atlas of Human Infectious Diseases (Wiley-Blackwell 2011)

To create the Atlas of Human Infectious Diseases, we gathered data on communicable diseases from either existing datasets, or, if unavailable, we created our own databases by reviewing the literature and other sources. We included drivers of infectious diseases in our mapping exercises, explaining the distribution of infections, such as global connectivity and sanitation. We also combined disease maps with relevant climatic variables like rainfall, similar to the approach taken for Lassa fever and histoplasmosis.
This was a very rewarding project with support from over 120 experts around the world, connected through the world-wide web, like a crowdsourcing project. Our talented Vietnamese mapping team provided excellent capacity to process this massive and valuable data. Through this work, I realised how important it is to have access to high-quality data and the value of professionals who can process this data robustly. A few years after completing the Atlas, we realised how drastically the world of infectious diseases had changed. Many of the disease maps we made just a few years ago already required a thorough update. Clearly, we need a better understanding of the many infectious diseases out there and the conditions of transmission, to make better predictions about where diseases occur or may occur in the future. To diagnose an outbreak and to predict its progression requires good diagnostic data.
Clinical microbiology
Good clinical microbiology facilitates direct patient care, feeds surveillance databases for public health purposes and is critical for investigating outbreaks.
Reliably detecting or excluding the microbiological cause of an infectious disease is central to clinical microbiology. For this purpose, several direct and indirect methods have been developed. Microscopy is a direct method used to visualise fungi, parasites like malaria, or bacteria. We have other techniques, such as culturing organisms in combination with mass spectrometry or detecting the pathogen’s specific genes through polymerase chain reaction (better known as PCR) or detection using a sequencing approach. There are also indirect methods that can identify a specific host antibody response to the pathogen using serological techniques.
Because the epidemiology of infectious diseases are dynamic and outbreaks can occur suddenly, a clinical microbiologist needs to be flexible and must implement new diagnostic strategies if the circumstances demand so. Unfortunately, devastating epidemic diseases like Ebola and outbreaks of drug-resistant infections often originate in resource-limited settings with weak health systems, where laboratory diagnostics are scarce. This creates the perfect circumstances for infectious diseases to multiply and spread. In our highly-connected world, no country is an island – disease can potentially reach any country, rich or poor, nearby or far away. Controlling infectious diseases in low-resource settings directly contributes to controlling these diseases in high-income countries like the Netherlands.
Digitisation provides the means to scale up the transfer of knowledge and technologies used to diagnose and manage infectious diseases in these higher-risk, low-capacity settings. This is already happening with molecular and sequencing techniques, and also with the older microscopy and culturing techniques. The possibilities created by billions of people connected with access to strong computer processing power and to instant knowledge and skills are countless.
Smartphones are combined with microscopes to read and analyse blood slides for malaria without the need of a trained laboratory technician and can report the results to the attending healthcare worker. These self-reading microscopes are currently being evaluated in field trials. This is a beautiful example of how a 350-year old technique is being digitised and contribute to diagnosing malaria, tuberculosis, and other infectious diseases worldwide. The microscope was invented in the seventeenth century by Antoni van Leeuwenhoek, who lived in an age of important scientific discoveries.
I would like to briefly touch upon this piece of history and share with you an important message that is relevant to our times.
Antoni van Leeuwenhoek
When Antoni van Leeuwenhoek’s first wife died, he started making microscopes. He was 38 years old at the time. He could experiment without financial worries as he remarried a woman from a wealthy family. He became very successful in making microscopes, which could magnify up to 260‐fold. With his microscopes, he examined everything from pond water, wood, hair, and even the eyes of insects. Around 1676, in an effort to determine why peppers are so spicy, he started to experiment with ‘pepper water’.
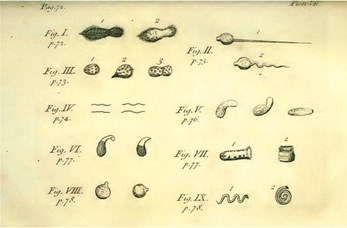
In figure 4 you can find the sketch he made from observing a sample from a bottle containing pepper water that turned cloudy. This drawing is believed to be the first sketch of bacteria. The birth of microbiology...
This revolutionary discovery cannot solely be ascribed to Van Leeuwenhoek’s achievements, driven by his tireless work ethic, his exceptional lens making skills and his observational skills. He was also the right man at the right time in the right country. He was living at the start of the Enlightenment, a time of paradigm shifts and important scientific discoveries that shaped the world by people like Newton, Spinoza, Descartes and others. Like Antoni van Leeuwenhoek, we now also live in an age of transformative scientific achievements. And we have a new lens, the genome sequencer, which allows us to observe and study natural life like never before, such as the human microbiota.
The human microbiota
The human body is colonised by a vast number of microbes. Collectively referred to as the human microbiota, it has a mutualistic relationship with the human host. The human microbiota has been jokingly called the organ system for infectious disease specialists or clinical microbiologists, just like the neurologist has the nervous system and the ophthalmologist the eye. In addition to their role in digesting food, these microbes can shape your immune system, keep potential pathogens at bay, play a role in how you feel and many other things.
The human microbiota is dynamic and changes with age, disease states, diet and other factors. And we are continuously sharing these microbes when we interact with each other, for example through a simple handshake. Inside of us, these microbes also interact with each other: communicating and sharing genes.
Dysbiosis of the human microbiota has been linked to many health issues, ranging from asthma to intestinal disorders like irritable bowel syndrome – not the usual areas of work for clinical microbiologists. Well-known pathogens are also among our microbiota, which do not usually cause us harm as the peer pressure of the other bacteria requires them to behave. However, a change in the microbiota can result in these opportunistic pathogens causing diseases. Clostridium difficile, which leads to severe diarrhoea, is a well-known example of this.
We do not yet understand how microbiome data can be used to improve patient management or to prevent disease; however, several studies have shown that, for instance, faecal transplantation with a diverse microbiome can improve health. Most clinical microbiologists are not used to analysing communities of microbes like the microbiota, an area in which environmental microbiologist feel more at home. This is a pivotal moment as medical microbiology and environmental microbiology are getting ever closer to each other. I advise residents in clinical microbiology and others to seize this opportunity and to follow the pioneering spirit of Antoni van Leeuwenhoek. Instead of using an optical lens, you will wield the powerful sequencing lens to study microbial life in relation to disease.
The power of digitisation
To process the ever-increasing volume of information, including big sequence data sets combined with patient characteristics from electronic patient files, we need more and more computer power, which will soon reach the capacity of the human brain and eventually the combined power of all human brains on this world and more. The speed of current breakthroughs is unprecedented and evolves at an exponential pace. A milestone was reached last year when artificial intelligence (AI) beat a master in the very complex Asian game ‘Go’. AI allows computers to learn and improve by analysing lots of examples instead of being programmed. We can incorporate artificial intelligence into our daily work to give us more time to do what we are good at as humans: being creative, understanding context and making human connections.
AI is already disrupting the medical specialties of radiology and pathology, which in turn support specialties like clinical microbiology. Artificial intelligence is capable of interpreting scans better than humans, as it tirelessly learns from millions of scans, remains objective and can see information invisible to the human eye. It also keeps working and learning 24/7. The possibilities to improve the quality of our work and patient outcomes and simultaneously reduce costs are numerous, and I believe this potential should be embraced more. Clinical microbiology needs to plan strategically for a future in which artificial intelligence is part of the daily routine and helps to make us better at what we do. The Radboud REshape Center and Philips have started exploring these possibilities together.
Let me take you through the different steps of clinical microbiology to highlight where digitisation can make a difference.
Clinical microbiology is known for taking its time. To get results, we often have to wait for pathogens to grow enough for identification and drug-susceptibility testing. The faster we can secure test results, the higher the clinical impact. In order to see where we can improve and speed up our work, we need to untangle the diagnostic process: from deciding whether or not to test, to collecting and sending the right sample, to analysing it and finally to reporting it. Laboratory tests are ordered on a daily basis, even when the pre-test probability of the patient with the disease is very low and a laboratory test would not have any added value and may even cause harm. As most hospitals today have electronic patient files, we have the potential to reduce unnecessary testing and increase required testing by matching clinical characteristics with test ordering. After ordering a test and collecting the sample, we need to ensure that it arrives in a timely fashion at the correct laboratory.
Diagnostic errors account for up to 10 percent of patient deaths.1 The majority of errors, around 60 percent, happen in the pre-analytic phase, which include the incorrect ordering of tests, the mislabelling of specimens or specimens getting lost.2 In case of mislabelling, a patient may get the wrong diagnosis or the wrong treatment, with a prolonged disease, disability or even death as an outcome. By mid-2017, Radboudumc will have a central specimen receiving area: a unique collaboration between the different Radboudumc laboratories. Nurses and transport staff only need to go to one desk rather than seven different places throughout the hospital, thereby reducing confusion and errors.
We are also considering the option of using radio frequency identification in specimen labels to allow us to track specimens throughout the hospital. I can track my Amazon book order at any time, but not a precious specimen needed for an accurate diagnosis? In addition to specimens, anything critically important can be labelled with radiofrequency tags (from healthcare workers to patients to hospital beds), which will enhance both diagnostics and infection control efforts.
Now that the sample has arrived in the clinical microbiology laboratory, the actual testing can be done; here, too, the techniques are changing.
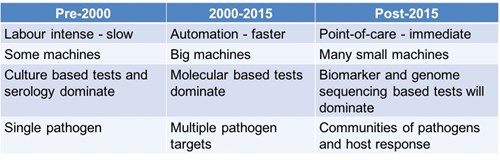
Before 2000, we had limited methods, such as cultures and serology, to detect a limited number of pathogens. This was also very labour-intensive and time-consuming. Over the past fifteen years, we have automated more and integrated molecular diagnostics into routine clinical microbiology, increasing the speed and also the number of pathogens that can be simultaneously identified. The drawback of molecular-based tests like PCR is that you can only find what you are specifically looking for. Furthermore, some pathogens may not have the target used in the test or may even ‘lose’ the target, like we have seen with malaria rapid tests.3 Pathogens lacking or losing the target have a survival advantage because once a pathogen is detected, both the doctor and the patient prefer to destroy it. As a result, a pathogen without a target may become dominant once a test is scaled up – this is microbial evolution driven by diagnostic testing! Fascinating.
It is the role of clinical microbiologists to monitor whether diagnostic tests remain sufficiently sensitive and accurate.
Considering our aging population and rising healthcare costs, we need to develop and evaluate diagnostic strategies that improve health and also reduce healthcare costs. Fast and flexible point-of-care systems close to the patient have a lot of potential in any setting. In addition, sequencing technology is expected to one day replace most culture-based techniques, but many years will pass before we reach the required standards. Another development is testing the host response with point-of-care biomarker tests, which can guide the attending doctor to the best empiric treatment and the most useful testing.
Whole genome sequencing of single pathogens, which is different from the sequencing approach of the microbiota I mentioned earlier, has already become part of the routine diagnostic arsenal of an academic clinical microbiology laboratory, made possible by affordable sequencing technologies. It is used for outbreak management, but also has the potential to support treatment decisions. In 2016, we strengthened our molecular team with bioinformatics expertise to set up a whole genome sequencing pipeline in collaboration with the genetics department and the Centre for Molecular and Biomolecular Informatics (CMBI). For Mycobacterium tuberculosis we will develop a separate WGS pipeline to genotype TB and detect drug resistance, which is normally a lengthy process. This will help to give the right treatment to these very sick patients much sooner.
WGS costs will decrease further once implemented on a larger diagnostic scale, meaning less-skilled staff will suffice over time. Placing bench-top sequencers in microbiology laboratories, including those in low-resource settings, combined with a fully-automated workflow and analysis may completely change the landscape of controlling infectious diseases like TB worldwide.4
Now I would like to discuss the issue of antibiotic resistance.
The tide of antibiotic resistance
Despite all of our digital advances, hospitals worldwide have different computer systems for direct patient care and laboratory management. The majority of the available information is underutilised, despite the fact that we need to provide more data driven care. Are patients, for instance, getting antibiotics that do not match what we test in the lab?
In close collaboration with the Information Management team, the Radboudumc Antibiotic Stewardship-team built algorithms to monitor antimicrobial treatment in our hospital, to determine whether it is appropriate and to provide advice if needed. It is also cost-effective, as we can switch patients from intravenous to oral medication sooner. More IT advancements can be made by sharing and analysing infectious disease data on a regional level, particularly to control the spread of antibiotic resistance. This year, the Dutch Ministry of Health will start financially supporting ten health networks (Dutch: zorgnetwerken) to better coordinate the response to the threat of antibiotic resistance.
In the province of Gelderland, key stakeholders formed the Gelders Antibiotic resistance and Infection prevention Network (GAIN). Considering the changing healthcare landscape, with more care being provided outside the hospitals, these networks are critical in monitoring healthcare-related infections and drug resistance and improving infection control and antibiotic stewardship in a non-hospital setting. We also contribute to other initiatives regionally, nationally and globally, such as our cross-border initiatives with Germany. These types of regional collaborations are important tools for controlling antibiotic resistance. The Netherlands are exceptionally good at that.
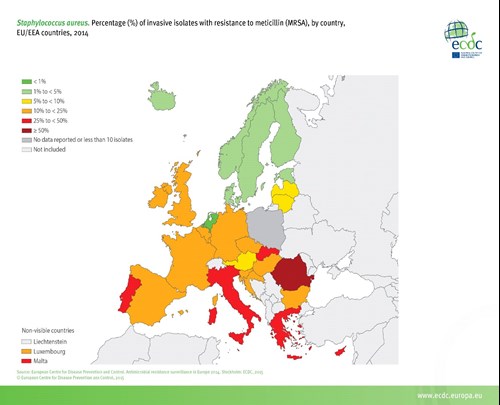
The Netherlands has the lowest rate of methicillin-resistant Staphylococcus aureus in the world, but also relatively low resistance rates to many other pathogens (see figure 5). We achieved this great success with a bundle of measures rolled out nationally. The low resistance rates we experience are being jeopardised by the rising antimicrobial resistance rates around us, reaching the country through travel and trade. One-third of healthy people who are negative for drug resistance before travel acquire resistant bacteria during international travel, with the highest number of acquisitions among those visiting Asia.5
As you are all aware, the Netherlands is below sea level and to protect us from the sea we invested in the Delta Works. Now we are facing the tide of drug resistance. The national policies we have and the recently implemented health networks can be viewed as the Delta Works for antimicrobial resistance. Instead of building stronger dykes, we need to slow the rise of drug resistance or preferably lower it. We can achieve this by taking our expertise and research activities to the areas where drug resistance is abundant, like Asia. That is where I want to contribute.
In September 2016, the UN Assembly agreed to act on antimicrobial resistance. For the first time, heads of state committed to taking a broad, coordinated approach to address the root causes of antimicrobial resistance across multiple sectors, especially human health, animal health and agriculture. This is just the fourth time a health issue has been taken up by the UN General Assembly (the others were HIV, non-communicable diseases and Ebola).
The deadly consequence of antimicrobial resistance and the global nature of this threat are illustrated by the recent death of an American woman due to a resistant infection acquired in India. No antibiotic available in the US was able to save her.
Hotspot Asia: treatment practices in Vietnam
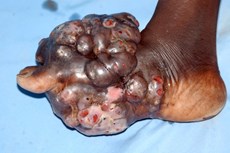
I had a comparable experience with a patient who was admitted to the ICU of a Vietnamese hospital. The young woman was recovering from her illness but developed ventilator associated pneumonia due to a multi-drug resistant bacterium, only sensitive to an antibiotic known as polymixin or better known as colistin, an antibiotic abandoned many years ago for systemic treatment due to its toxicity. The drug was not available for human use in Vietnam and the patient died. Five hundred meters away from the hospital was a veterinary pharmacy with buckets of this drug, which is usually mixed with animal feed, illustrating the One Health and complex nature of this problem. Systemic colistin is increasingly used worldwide despite its toxicities due to the rising number of drug resistant infections. Colistin is now part of the empiric treatment provided by many intensive care units in Vietnam, treating patients suspected of having a hospital-acquired infection, of which most are now carbapenem (last resort antibiotic) resistant.
Multi-drug resistance is rampant in Asian hospitals, especially in intensive care units where there is inadequate infection control.
Where to start and improve the situation?
The issue of drug resistance reflects a health system failure, or more precisely, a ONE HEALTH system failure. LMIC countries thus far have relied on antibiotics to control infectious diseases rather than prevent them with proper infection control. With current high resistance levels, they need to ramp up their prevention efforts. Surveillance of drug resistance is a basic measure to make informed decisions about the next steps. We are involved in several initiatives to implement effective surveillance strategies. As antibiotics are the drivers of resistance, we need to ensure that those who need it, get it and those who do not need it, do not get it.
Ninety percent of antibiotics in Vietnam are dispensed without a prescription, even though the law requires one. To find potential control strategies we need a better grasp on our understanding of how antibiotics are used in the community setting. We recently launched a study with Oxford and the INDEPTH Network the ABACUS aimed at identifying the determinants of appropriate antibiotic use in low and middle income countries to identify potential interventions to address antibiotic misuse (http://www.indepth-network.org/projects/abacus). However, we also need to act now and evaluate promising existing interventions. In the community setting, most antibiotic overuse is by patients with acute respiratory tract infections, which are often self‐limiting and do not require antibiotic treatment.
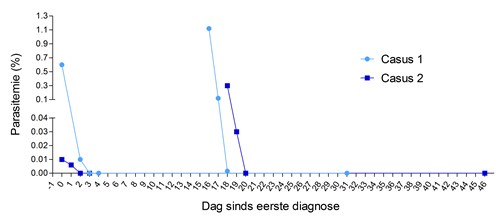
A major reason for uncontrolled antibiotic use for respiratory infections is diagnostic uncertainty. In low‐income settings, physicians worry about their patients’ inability to access healthcare if their condition deteriorates. These factors motivate overuse of antibiotics. We found that C‐Reactive Protein or CRP point-of-care testing for respiratory infections can safely bring down antibiotic overuse in Vietnam.6 CRP is an inflammatory protein in the blood. A low test result means that no serious infection is present and no antibiotics are needed.
In Vietnam, CRP testing has the potential to avert the prescription of at least 1.3 million courses of antibiotics every year in primary care patients, and this is a conservative estimate. The success of this trial prompted us to evaluate it in other low-income countries. CRP can be made for less than a dollar per test and can be combined with rapid tests for malaria or dengue fever – a huge potential for reducing AB use globally. I look forward to working on these scale-up studies and improving the correct use of antibiotics.
In addition to improving antibiotic use, we need to understand the reservoir of resistance and hotspots for transmission. If Dutch travellers can easily pick up drug-resistant bacteria in Asia, what about locals?
Asia is a hotspot for drug resistance to commonly-used antibiotics critical for healthcare, and recently even to the last-resort antibiotic colistin. Besides the overuse of antibiotics, poor access to clean water and sanitation contribute to the spread of resistance. People defecate in the same water they use to wash clothes and cook with.
In a household cohort in northern Vietnam, we regularly sampled the stool of household members, their animals, the water they use and the food they eat. Preliminary findings show high rates of resistance to common antibiotics and also to colistin in healthy humans and dogs.
As already mentioned, in Asia, colistin is commonly used as prophylaxis in agriculture. The resistance gene for colistin, called MCR‐1, is positioned on a plasmid and can easily be shared between bacteria. This plasmid with MCR‐1 emerged recently and can spread very easily. It was present in 80% of the dogs and humans we tested. Bacteria excel at sharing useful genes including resistance genes: the sharing economy was already invented by nature. The colistin resistance that originated in Asia is a potential threat to Dutch intensive care patients who receive colistin as part of prophylaxis. Antibiotic prophylaxis in general is critical for protecting patients from acquiring an infection during a medical intervention. Antibiotic resistance is increasingly causing trouble here, requiring targeted solutions, both abroad and also at Radboudumc.
With the Radboudumc urology department we will evaluate whether culture-guided antibiotic prophylaxis for prostate biopsies reduces the number of post-biopsy infections. We increasingly observe breakthrough infections with ciprofloxacin-resistant bacteria after a prostate biopsy, and ciprofloxacin is the drug we use for prophylaxis. The strategy is to test patients before biopsy for the presence of resistant bacteria in the rectum and, in case of ciprofloxacin resistance, an alternative antibiotic will be used as prophylaxis. We look forward to conducting this diagnostic clinical trial, which will have an impact on guidelines for this widely-used intervention and potentially also on other surgical interventions where culture-guided prophylaxis may be beneficial.
Outlook
We are experiencing breath-taking advances in computation, artificial intelligence and physical and biological sciences, also known as the Fourth Industrial Revolution. It is predicted that the USA within will see a $9 trillion reduction in employment costs due to AI over the next decade, including knowledge work in healthcare. The McKinsey Global Institute states that AI contributes to a transformation of society ‘happening at roughly 3,000 times the impact’ of the Industrial Revolution. Impressive!
Is clinical microbiology sufficiently prepared and able to adapt to this reality and seize the opportunity to improve prevention, diagnosis and treatment of infectious diseases around the world? I think that we have made a cautious start but we can be bolder. Like Antoni van Leeuwenhoek before, this is our carpe diem moment.
Billions of people are connected and communicating through smartphones. This has created a society where people reside in their own digital echo chambers. Opinions – not facts – spread easily: just look at the current developments regarding the effectiveness and risks of vaccination. This is a reality that we need to consider carefully. As academics, we bear the collective responsibility to bring forward facts and to guide the greater public debate.
We still have a lot of work to do!
Closing remarks and acknowledgements
Radboud University successfully focuses on infectious diseases; however, we are still too fragmented and want to unite our efforts more effectively in the coming months as part of the Radboud Center for Infectious Diseases (RCI). And we will undertake this effort with many partners, both close and far away. Radboud University provides a unique opportunity to address the global issue of antimicrobial resistance with a multidisciplinary approach, involving social sciences, pharmacology, computer sciences, medicine, international law and environmental microbiology. With full optimism we are building the necessary bridges to deal with small bugs and their big data.
Kohn LT, Corrigan JM, Donaldson MS. To Err Is Human: Building a Safer Health System. Washington, DC: National Academy Press; 1999.
Carraro P, Plebani M. Errors in a stat laboratory: types and frequencies 10 years later. Clin Chem. 2007;53:1338-42.
Kozycki CT, Umulisa N, Rulisa S, Mwikarago EI, Musabyimana JP, Habimana JP, Karema C, Krogstad DJ. False-negative malaria rapid diagnostic tests in Rwanda: impact of Plasmodium falciparum isolates lacking hrp2 and declining malaria transmission. Malar J. 2017 20;16:123. doi: 10.1186/s12936-017-1768-1.
Votintseva AA, Bradley P, Pankhurst L, et al. Same-day diagnostic and surveillance data for tuberculosis via whole genome sequencing of direct respiratory samples. J Clin Microbiol. 2017 Mar 8. pii: JCM.02483-16. doi: 10.1128/JCM.02483-16.
Arcilla MS, van Hattem JM, Haverkate, et al. Import and spread of extended-spectrum β-lactamase-producing Enterobacteriaceae by international travellers (COMBAT study): a prospective, multicentre cohort study. Lancet Infect Dis. 2017;17:78-85. doi: 10.1016/S1473-3099(16)30319-X.
Do NT, Ta NT, Tran NT, et al. Point-of-care C-reactive protein testing to reduce inappropriate use of antibiotics for non-severe acute respiratory infections in Vietnamese primary health care: a randomised controlled trial. Lancet Glob Health. 2016;4:e633-41. doi: 10.1016/S2214-109X(16)30142-5.